Many countries such as Sweden rely on nuclear power to meet their energy needs. As a result, various types of deep geologic repository concepts for the disposal of nuclear waste are being planned worldwide. The goal of this PhD project is to study radiation-induced chemistry at the molecular level in the context of nuclear waste disposal, using advanced theoretical and experimental techniques, such as molecular simulations, gamma-radiation and X-ray synchrotron-based methods. This 4-year PhD project is funded by the Swedish Radiation Authority (SSM), and builds upon prior research in the Holmboe group in collaboration with Prof. Mats Jonsson, Applied Phys. Chem, KTH, Sweden.
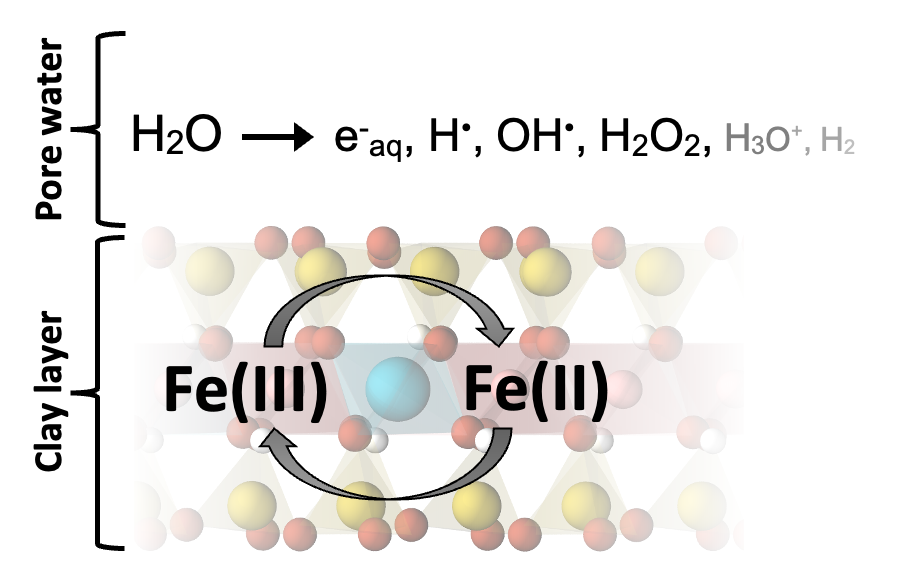

Figure 1. Schematic of flip-flopping of structural Fe(III) to Fe(II) and back, due to reactions with the primary radicals from water radiolysis.